In the Medical Cell Biology group, we generate and work with
human induced pluripotent stem cells (iPSCs)
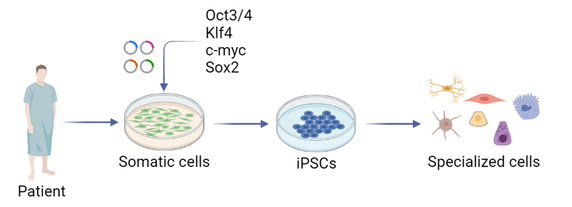
- Stem cells offer tremendous value to regenerative medicine. These cells renew indefinitely and can differentiate into almost any given cell type in the human body. Human embryonic stem cells have been employed for decades, but more recently iPSCs have revolutionized the field of regenerative medicine. These induced stem cells provide many advantages: they pose less ethical concerns and exhibit the genetic background of the donor patient. iPSCs are readily generated in our lab, using an integration-free reprogramming approach. Invasive sample collection from the patient is avoided by working with a simple urine sample. Exfoliated renal epithelial cells are isolated and expanded from the urine sample, serving as excellent starting material for reprogramming. The commercial Sendai Cytotune 2.0 kit (Life Technologies) is used for maximum efficiency and reliability.
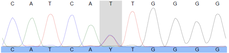
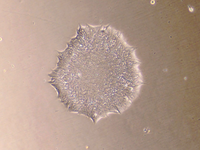
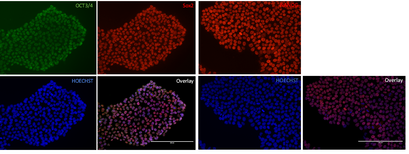
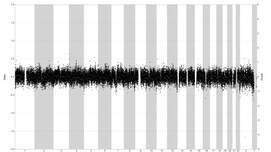
- The generated iPSCs undergo an extensive quality check. We check their morphology using microscopy, which should resemble round, tightly packed colonies of stem cells, with smooth edges and a transparent body. Pluripotency is confirmed by staining for specific factors, such as POU5F1, NANOG and SOX2. Shallow whole genome sequencing (SWGS) will alert us in case of genomic aberrations, such as deletions or duplications. The differentiation potential is confirmed by directing the iPSCs towards the three different germ lines, which are validated using PCR. The pathogenic variant of interest (if present) is checked using Sanger sequencing.
- With the rise of CRISPR genome editing, we can make precise changes to the genome. This allows us to correct pathogenic variants in the genome. In our lab, we use the following workflow, where we add a complexed Cas9, the protein that cuts the DNA with the single guide RNA, which recognizes the target site of interest. Alongside this RNP complex, we add a single stranded oligodeoxynucleotide repair template by means of nucleofection. When Cas9 introduces a double strand break, the cell will try to repair this break by a process called NHEJ (non-homologous end joining). This process however is prone to errors, introducing other mutations. By providing a repair template, the cell uses an alternative, more precise repair method: homology directed repair. After recovery, the cells are sorted one by one into a new well, to ensure monoclonality. Repair of the targeted pathogenic variant is evaluated, after which the successfully corrected clones are further expanded and subjected to the same characterization as described above. These generated lines serve as isogenic controls, with an identical genetic background (apart from the corrected pathogenic variant) as to the diseased iPSC line.
In the Medical Cell Biology group, we generate and work with bovine embryonic stem cells
- Bovine embryonic stem cells (bESCs) are becoming increasingly important in both regenerative medicine and the production of cultured meat. Like iPSCs, bESCs possess the ability to renew indefinitely and differentiate into almost any cell type. This unlimited proliferation potential makes them invaluable for various applications. Derived from early-stage embryos, bESCs share the same versatile characteristics as human iPSCs, providing a robust platform for studying developmental processes and tissue regeneration in bovine species. In the context of cultured meat, bESCs offer a promising avenue for producing high-quality, structured meat by differentiating into muscle cells and other tissue types. Their use in research continues to advance our understanding of cell differentiation and tissue regeneration, paving the way for innovative therapies, veterinary treatments, and sustainable food production solutions.
- The generated ESCs undergo an extensive quality check. We check their morphology using microscopy, which should resemble round, tightly packed colonies of stem cells, with smooth edges and a transparent body. Pluripotency is confirmed by staining for specific factors, such as POU5F1, NANOG and SOX2. Shallow whole genome sequencing (SWGS) will alert us in case of genomic aberrations, such as deletions or duplications. The differentiation potential is confirmed by directing the ESCs towards the three different germ lines, which are validated using immunology and qPCR.
- In our lab, we focus on improving muscle maturation, hypertrophy, and tissue visco-elastic properties through stimulation experiments on muscle-derived myoblasts. We employ pulsed-electromagnetic field stimulation (PEMFS) and mechanical stimulation (MS), methods adapted from human tissue-engineered skeletal muscle constructs. For cultured meat production, we use either bESCs or bovine myoblasts to differentiate into muscle cells. PEMFS involves short-interval stimulations to enhance the fusion capacity of myoblasts. MS is applied on bio-artificial myoblast-containing muscle constructs (BAM) to promote metabolic activity and increased stiffness, as well as to encourage bovine myofiber formation. To visualize and process whole-mount bovine muscle constructs, we use optical clearing techniques such as Cubic clearing and 3DISCO clearing. These methods improve tissue transparency and facilitate deep antibody penetration, enhancing our ability to study and refine the muscle constructs.